SFTP Offline Data Ingestion
Overview
This document defines the SFTP directory structure, file formats, metadata requirements, configuration setup, and command-specific behaviors for offline data ingestion. It serves as a reference for preparing, uploading, and validating ingestion files in a consistent manner. The guidelines ensure reliable processing, error handling, and traceability across ingestion workflows.
It applies to the following ingestion types:
-
UPSBATCH: User profile and segment ingestion
-
UPSLINK: User identity linking
-
HIVEUPLOAD: Hive table uploads
SFTP Directory Structure
All ingestion files must be uploaded to the configured SFTP base path (ftpBasePath). The directory structure is organized to separate ingestion types and maintain a clear lifecycle of files from upload to processing and archival.
Directory Layout
/
└── offline_data/ (ftpBasePath – configurable)
├── upsbatch/ UPS batch ingestion
├── upslink/ User linking ingestion
├── hiveupload/ Hive table uploads
└── archive/
├── processing/ Temporary processing area
│ ├── upsbatch/
│ ├── upslink/
│ └── hiveupload/
├── processed/ Successfully processed files
│ └── YYYYMMDDHHMMSS/
│ ├── upsbatch/
│ ├── upslink/
│ └── hiveupload/
└── failed/ Failed files with error logs
└── YYYYMMDDHHMMSS/
├── upsbatch/
├── upslink/
└── hiveupload/Timestamped directories ensure traceability and auditability. Failed files are stored along with corresponding .error logs, which provide details about processing failures for debugging and reprocessing.
Supported Compressed File Formats
This section defines the allowed archive formats for uploading ingestion files. Restricting supported formats ensures compatibility with the ingestion pipeline and prevents processing errors due to unsupported compression types.
Configurable via:
sftp.allowed.archive.extensionsDefault Supported Formats:
-
.zip: Standard ZIP compression
-
.tar.gz: TAR archive with GZIP compression
Only these formats are accepted at the SFTP ingestion layer.
Supported Data File Formats (Inside Archives)
This section specifies the allowed data file formats contained within compressed archives. These formats are chosen to support structured and semi-structured data ingestion efficiently.
Configurable via:
sftp.allowed.file.extensionsDefault Supported Formats:
-
.csv: Comma-separated or custom-delimited values
-
.txt: Plain text
-
.json: JSON (JSON Lines format for batch data)
File Naming Convention
There are no strict constraints on file naming, allowing flexibility for different data providers and workflows. However, meaningful and consistent naming is recommended for easier identification and tracking of files. Any filename is accepted if the extension is valid.
Examples
-
segments_20240115.zip
-
user_linking_grouped_20240115.tar.gz
-
products_hive_upload.zip
Compressed File Content Requirements
Each compressed file must include both metadata and data files to ensure proper processing. The metadata file defines how the ingestion should be executed, while the data files contain the actual records to be processed.
Each compressed file must contain:
-
Metadata File (Required)
Exactly one of the following:
-
metadata.json
-
metadata.properties
-
metadata.txt
-
-
Data Files (Required)
-
One or more files with supported extensions (.csv, .txt, .json)
-
Files missing a valid metadata file will be rejected.
Configure SFTP Credentials and Site Mapping
Configure Credentials in Dashboard
SFTP credentials must be configured in Site Configuration RR within the Personalization Platfom. These credentials are used to authenticate and securely connect to the SFTP server for file ingestion.
Under the Omnichannel Site Configurations, create FTP Username and FTP password to establish the SFTP connection and process ingestion files.
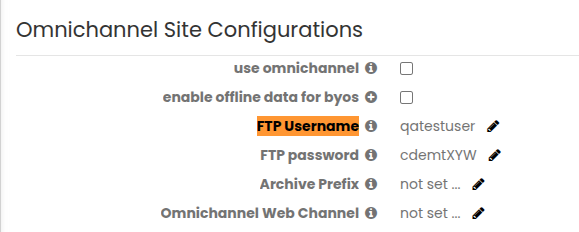
Update BuildFTP Configuration
After configuring credentials in the dashboard, update the BuildFTP configuration to map sites to their respective SFTP paths. This ensures that files are routed and processed correctly for each site.
Add/Update the following entries to config.properties:
sftp.allowed.sites=<allowed_sites_list>
sftp.ftp.base.paths.per.site=<site_id_1=path1;site_id_2=path2>Example
sftp.remote.server.host=ftp.richrelevance.com
sftp.allowed.sites=801,1218
sftp.ftp.base.paths.per.site=801=/path/offline;1218=/userpath/batchCommand Execution Reference
UPSBATCH Command Specification
The UPSBATCH command is used for ingesting user profiles, segments, and attributes. It supports both full and incremental data loads and allows optional optimizations such as columnar storage.
Supported Options
| Option | Mandatory | Default | Description |
|---|---|---|---|
| attribute | Yes | None |
Defines the attribute type identifier and determines how the data is processed. It must follow strict validation rules to ensure consistency. Allowed characters: alphanumeric, underscore (_), dash (-). Validation errors:
Examples
|
| loadtype | No | full |
Specifies whether the data should replace existing records or be applied incrementally. This allows flexibility based on the ingestion use case. Allowed values:
Any value other than delta is treated as full. |
| columnarSupport | No | false |
Enables optimized storage for supported attributes when specific conditions are met. This improves performance for large-scale data processing scenarios. Allowed Values: true, false Enabled only when:
When columnarSupport is not enabled, segment uploads using loadtype=delta override existing segment data. Only the segments present in the latest file are retained. When columnarSupport=true, previously uploaded segments are retained. New segments from subsequent uploads are appended instead of replacing existing data. This ensures cumulative segment building across multiple uploads. Use the following command to enable columnar support for segment uploads: upsbatch <filename> -attribute rr-segments -columnarSupport true |
Examples
Example 1: rr-segments Full Load (metadata.json)
File Structure
segments_full_20240115.zip
├── metadata.json [REQUIRED]
└── segments_data.json [Data file]SFTP Upload Path
/offline_data/upsbatch/segments_full_20240115.zipmetadata.json
{
"command": "upsbatch",
"attribute": "rr-segments",
"loadtype": "full"
}Note: You may use either a .json or a .properties metadata file.
Example 2: rr-userattributes Delta (metadata.properties)
File Structure
userattributes_delta_20240115.tar.gz
├── metadata.properties [REQUIRED]
└── user_attributes.json [Data file]metadata.properties
command=upsbatch
attribute=rr-userattributes
loadtype=deltaExample 3: wine-segments Delta (metadata.json)
File Structure
wine_segments_delta_20240115.zip
├── metadata.json [REQUIRED]
└── wine_segments.json [Data file]metadata.json
{
"command": "upsbatch",
"attribute": "wine-segments",
"loadtype": "delta"
}UPSLINK Command Specification (User Linking)
UPSLINK handles user identity linking by associating multiple identifiers with a single user. It supports both real-time and batch processing modes to handle different data volumes.
Supported Options
| Option | Mandatory | Default | Description |
|---|---|---|---|
| command | Yes | upslink |
Command identifier. |
| linktype | No | explicit |
Determines how linking is processed.
Invalid Value Error: "unknown 'linktype' argument. Known values are: 'explicit' and 'grouped'" |
| loadtype | No | full |
Controls whether linking replaces existing data or updates incrementally.
Non-delta values default to full.
|
Data Format
The data must be provided in a specific delimited format to ensure correct parsing. Each row represents a group of linked user identities.
Format: Ctrl+A (\u0001) delimited CSV
user1^Auser2^Auser3
user4^Auser5^Auser6Rules:
-
Minimum 2 users per line.
-
Lines with fewer users are ignored.
-
First user is primary; others are alternate identities.
Examples
Example 1: Explicit and Full Linking (Default)
File Structure
user_linking_explicit_20240115.zip
├── metadata.json
└── user_links.csvUpload Path
/offline_data/upslink/user_linking_explicit_20240115.zipmetadata.json
{
"command": "upslink"
}Example 2: Grouped and Full Linking (Large-Scale)
File Structure
user_linking_grouped_20240115.tar.gz
├── metadata.properties
└── user_links_grouped.csvmetadata.properties
command=upslink
linktype=groupedExample 3: Grouped Delta Linking (Small Files)
File Structure
user_linking_grouped_delta_20240115.tar.gz
├── metadata.json
└── user_links_delta.csvmetadata.json
{
"command": "upslink",
"linktype": "grouped",
"loadtype": "delta"
}HIVEUPLOAD Command Specification
HIVEUPLOAD is used for uploading files directly into Hive tables. It allows flexible file naming while relying on metadata to define the target table.
Metadata Mapping
| FTP Option | Metadata Key |
|---|---|
| site hiveupload | command=hiveupload |
| -table <tablename> | table= |
Key Rules
These rules ensure consistent behavior during Hive ingestion and avoid ambiguity in table mapping.
-
Data file name does not need to match the table name.
-
Archive name can be arbitrary.
-
Table name is defined only in metadata.
-
All uploads use the default merchant database.
Example
File Structure
userfavorites_upload_20240115.zip
├── metadata.json [REQUIRED]
└── userfavorites.csv [Data file]Upload Path
/offline_data/hiveupload/userfavorites_upload_20240115.zipmetadata.json
{
"command": "hiveupload",
"table": "userfavorites"
}metadata.properties (Equivalent)
command=hiveupload
table=userfavoritesOnly one metadata file is required per archive.